- INTEGRAL EDITOR MESTRENOVA MANUAL
- INTEGRAL EDITOR MESTRENOVA FULL
- INTEGRAL EDITOR MESTRENOVA SOFTWARE
Everything is customisable to fit the desired format. Making publication quality images is very easy in MNova.
INTEGRAL EDITOR MESTRENOVA FULL
Use the right clicking followed by Properties allows full customisation of the overlaid or stacked spectra. To stack or overlay spectra, just select the spectra in the pages list, and click the relevant button. Any assignments made on the in 1D spectrum appear in the 2D to help with assignment on the other axis. To add a 1D spectrum to the axis of a 2D, you just drag and drop one from the pages list in the document to the desired axis. Processing 2D spectra in MNova is also very intuitive.
INTEGRAL EDITOR MESTRENOVA MANUAL
MNova struggles with automatic assignment in larger molecules, where a fair amount of manual correction will be required. MNova can report the multiplet analysis in journal or patent format for you to copy into your experimental. There are plenty of tools in MNova that can be used to manually edit the automatic assignments for complex or overlapping peaks. It’s worth checking through the automatic assignments manually, I’ve noticed it finding 13C peaks in the background noise fairly regularly. Automatic multiplet analysis followed by automatic peak assignment took only a minute or two. The drawing tool is quite clunky, so it may be easier to just paste structures from Chemdraw. Drawing the structure was the most time-consuming part of the analysis. The above screenshot of MNova took only a few minutes to make. The automatic analysis and assignment function is very good for small molecules. The whole process of manual analysis is intuitive and almost everything is customisable. Properties can be set as default or saved, so all future spectra opened will have the same format. Right clicking on the spectrum background and selecting Properties opens up a menu to change the appearance of the spectrum for everything from the size of the text to the thickness of lines. compound, impurity or solvent – distinguished by the colour of the label). Right clicking on an element allows editing, such as changing the integral value or the peak type (e.g. Elements of the spectra, such as peak shifts and integration values, can be moved around by simply clicking on them and dragging them to the desired position. Manual integration and pick picking can be quickly completed by clicking the relevant button or pressing the correct hotkey, followed by clicking and dragging over the correct area in the main viewer. Getting from the raw data FID to a readable spectrum takes only a few seconds. Clicking the Reference button (or pressing L) allows referencing of the spectrum using the solvent peak. The automatic phase correction isn’t perfect and can be corrected using the manual correction button (or shift P). These can be set to be performed automatically via Edit > Preferences > NMR. The top bar contains buttons for automatic phase correction and baseline correction. MNova will automatically process and Fourier transform the data according to the default processing parameters (these can be changed via Processing > Processing parameters). Muliple data files can be added at once and each will be a separate page within the document. ser file of the raw data into the MNova window. Raw data files can be added via File > Open, or by simply dragging and dropping the. The pages list and toolbars can be changed to floating menus if you would prefer to have more space in the window for the selected data set. The left-hand bar contains the tools for 2D and stacked spectra. Pressing some shortcut keys multiple times cycles through the possible functions (pressing Z multiple times changes between horizontal, vertical and box zoom). increasing and decreasing intensity is also completed by mouse wheel). Hovering over each button shows its function, and each button has shortcuts associated with them (e.g. The main bar above the display contains the tools for processing and analysing spectrum, as well as tools for drawing structures onto the spectra. The pages list can be used to browse different spectra within one document and 1D and 2D spectra can be included in the same document. Each spectrum has its own page (similar to a Powerpoint slide), with the currently selected data set filling the majority of the window. The default view resembles Powerpoint, making it very easy to start using.
INTEGRAL EDITOR MESTRENOVA SOFTWARE
The software can be downloaded from Mestrelab’s website (45-day free trial licences are available).
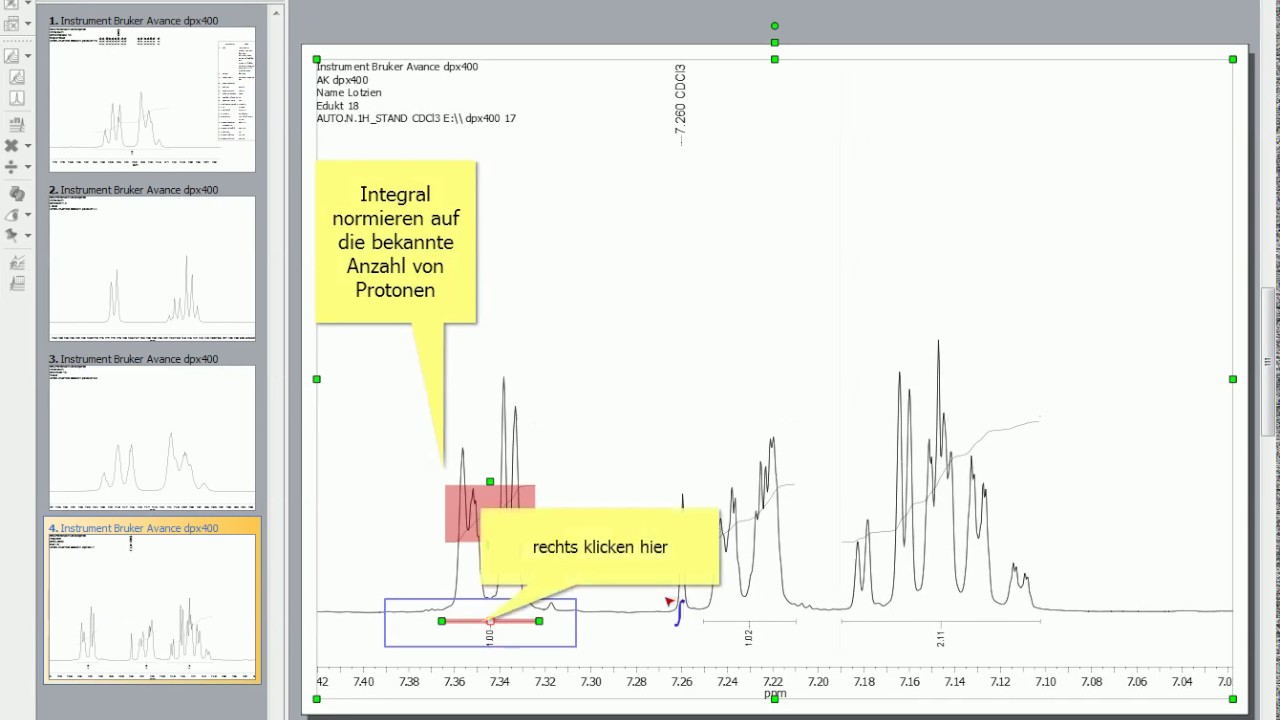
MNova NMR is Mestrelab Research’s NMR analysis program that can be used to quickly view, process and analyse both 1D and 2D spectra, as well as to easily produce publication quality assignments and images. A Review of MNova NMR (MestReNova 10.0.1 for Mac, 2015)
